BacterialTyper Documentation¶
- Version:
0.6.5
- Date:
Nov 20, 2023
Introduction¶
BacterialTyper is a pipeline that integrates multiple bioinformatic tools for the analysis of bacterial
whole genome sequence (WGS) data from isolated microbial cultured colonies. It generates genotyping information
and facilitates the interpretation of results.
The pipeline is written in Python with a modular architecture and based on open-source software and databases engines. The design of this bioinformatic tool allows comparing samples with an internal database (previously identified samples) and external databases.
- Multiple tasks are performed by several modules including:
preparation of raw data
generation of a virulence and resistance profile
bacterial strain identification
clustering based on sequence similarity
assembly and annotation
mobile genetic elements identification (plasmids, putative pathogenicity islands or phage insertions regions)
phylogenetic analysis
integration of metadata
preparation of results for integrative visualization
The tool uses and updates periodically external databases from different sources. It also allows the comparison of obtained results with those previously generated (internal database).
The BacterialTyper documentation includes:
A User’s Guide to get started.
An example Tutorial.
A list of Frequently Asked Questions (FAQs)
Some developer Guidelines to contribute to the project.
Additional information of the
BacterialTyperproject.A list of Glossary terms.
A list of Bibliography
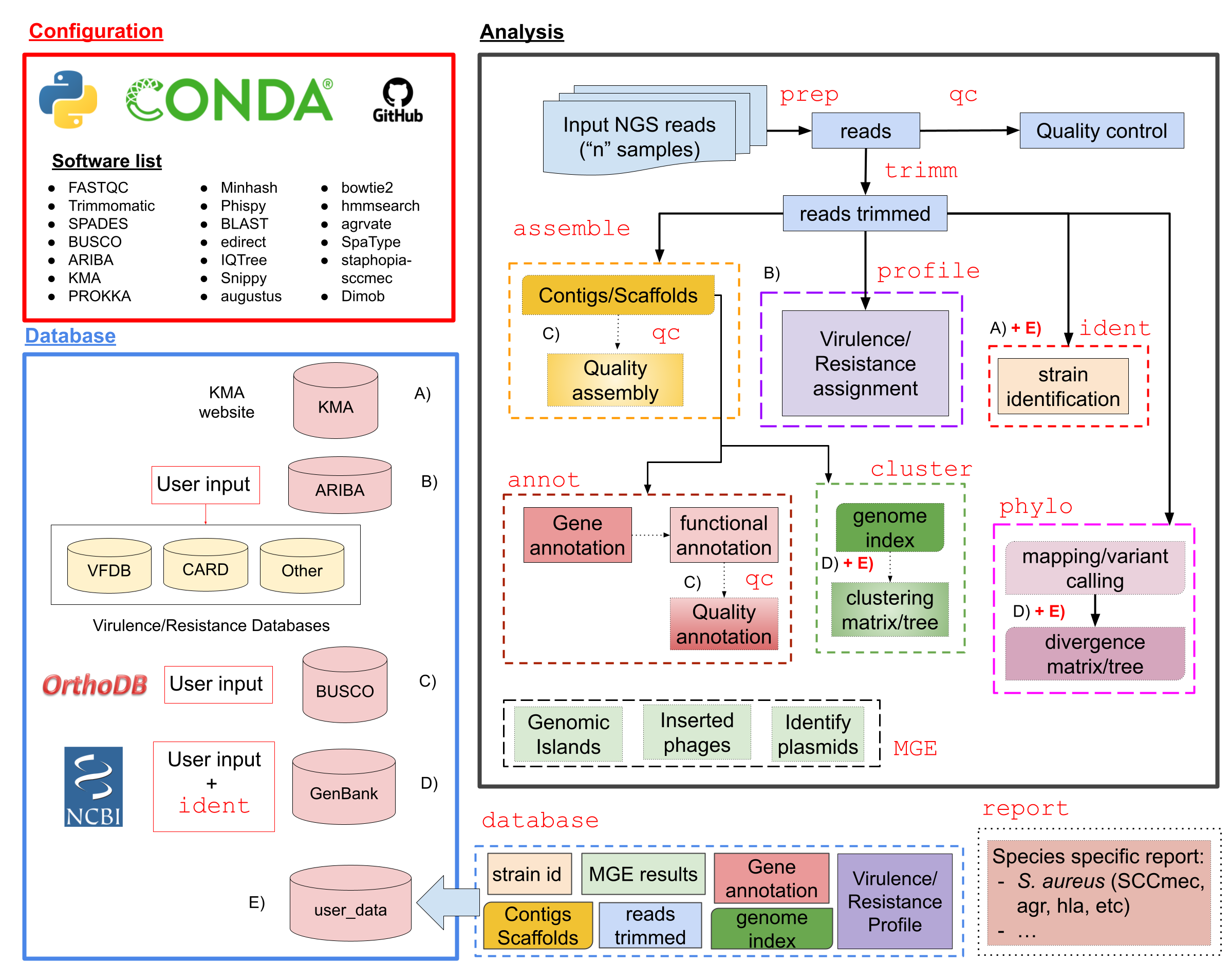
Pipeline Scheme¶
Here we show the scheme of the BacterialTyper bioinformatic tool. It is divided in four main actions:
Set the database: Several databases are downloaded from different websites/sources. They are automatically downloaded, indexed and/or updated after several days/months.
Identification analysis: for each sample of interest several bioinformatic steps are performed.
Populate the database: using the information generated in the identification analysis, user database is populated for later analysis.
Outbreak progression analysis: Using information in the database and samples of interest, user guides the analysis of the emergence or progression of an outbreak.